braindecode.models.InterpolatedBIOT#
- class braindecode.models.InterpolatedBIOT(chs_info, n_outputs=None, n_times=None, input_window_seconds=None, sfreq=None, n_chans=None, interpolation_method='spline', interpolation_mode='name_match', trainable=False, **kwargs)[source]#
Channel-interpolating wrapper around
BIOT.Channel
Accepts arbitrary user
chs_infoand projects them to the backbone’s canonical channel set viaChannelInterpolationLayer.For all other parameters and behavior see the backbone documentation reproduced below.
BIOT from Yang et al (2023) [Yang2023]
Foundation Model
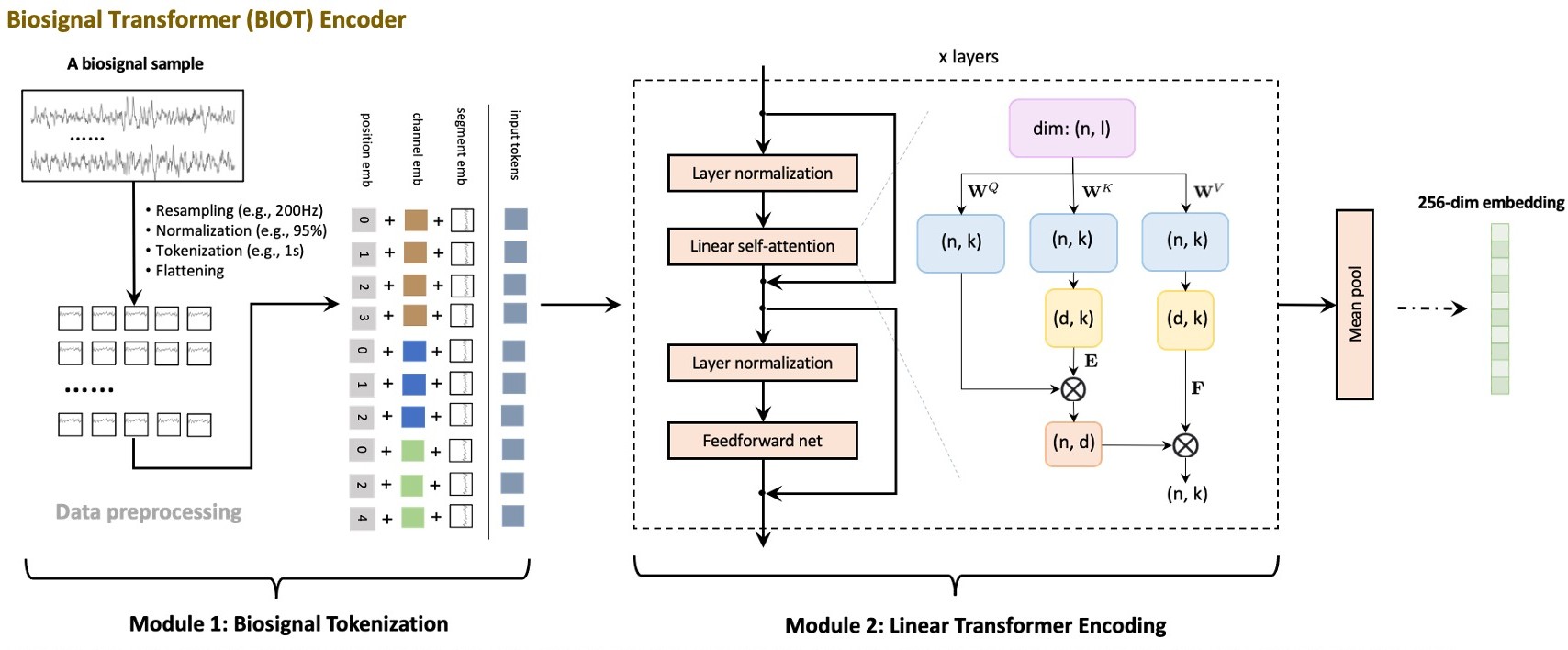
BIOT: Cross-data Biosignal Learning in the Wild.
BIOT is a foundation model for biosignal classification. It is a wrapper around the BIOTEncoder and ClassificationHead modules.
It is designed for N-dimensional biosignal data such as EEG, ECG, etc. The method was proposed by Yang et al. [Yang2023] and the code is available at [Code2023]
The model is trained with a contrastive loss on large EEG datasets TUH Abnormal EEG Corpus with 400K samples and Sleep Heart Health Study 5M. Here, we only provide the model architecture, not the pre-trained weights or contrastive loss training.
The architecture is based on the LinearAttentionTransformer and PatchFrequencyEmbedding modules. The BIOTEncoder is a transformer that takes the input data and outputs a fixed-size representation of the input data. More details are present in the BIOTEncoder class.
The ClassificationHead is an ELU activation layer, followed by a simple linear layer that takes the output of the BIOTEncoder and outputs the classification probabilities.
Important
Pre-trained Weights Available
This model has pre-trained weights available on the Hugging Face Hub. You can load them using:
from braindecode.models import BIOT # Load the original pre-trained model from Hugging Face Hub # For 16-channel models: model = BIOT.from_pretrained("braindecode/biot-pretrained-prest-16chs") # For 18-channel models: model = BIOT.from_pretrained("braindecode/biot-pretrained-shhs-prest-18chs") model = BIOT.from_pretrained("braindecode/biot-pretrained-six-datasets-18chs")
To push your own trained model to the Hub:
# After training your model model.push_to_hub( repo_id="username/my-biot-model", commit_message="Upload trained BIOT model" )
Requires installing
braindecode[hug]for Hub integration.Added in version 0.9.
- Parameters:
embed_dim (int, optional) – The size of the embedding layer, by default 256
num_heads (int, optional) – The number of attention heads, by default 8
num_layers (int, optional) – The number of transformer layers, by default 4
sfreq (int, optional) – The sfreq parameter for the encoder. The default is 200
hop_length (int, optional) – The hop length for the torch.stft transformation in the encoder. The default is 100.
return_feature (bool, optional) – Changing the output for the neural network. Default is single tensor when return_feature is True, return embedding space too. Default is False.
n_outputs (int) – Number of outputs of the model. This is the number of classes in the case of classification.
n_chans (int) – Number of EEG channels.
chs_info (list of dict) – Information about each individual EEG channel. This should be filled with
info["chs"]. Refer tomne.Infofor more details.n_times (int) – Number of time samples of the input window.
input_window_seconds (float) – Length of the input window in seconds.
activation (nn.Module, default=nn.ELU) – Activation function class to apply. Should be a PyTorch activation module class like
nn.ReLUornn.ELU. Default isnn.ELU.drop_prob – The description is missing.
max_seq_len – The description is missing.
att_drop_prob – The description is missing.
att_layer_drop_prob – The description is missing.
- Raises:
ValueError – If some input signal-related parameters are not specified: and can not be inferred.
Notes
If some input signal-related parameters are not specified, there will be an attempt to infer them from the other parameters.
References
[Yang2023] (1,2)Yang, C., Westover, M.B. and Sun, J., 2023, November. BIOT: Biosignal Transformer for Cross-data Learning in the Wild. In Thirty-seventh Conference on Neural Information Processing Systems, NeurIPS.
[Code2023]Yang, C., Westover, M.B. and Sun, J., 2023. BIOT Biosignal Transformer for Cross-data Learning in the Wild. GitHub ycq091044/BIOT (accessed 2024-02-13)
Hugging Face Hub integration
When the optional
huggingface_hubpackage is installed, all models automatically gain the ability to be pushed to and loaded from the Hugging Face Hub. Install with:pip install braindecode[hub]
Pushing a model to the Hub:
from braindecode.models import BIOT # Train your model model = BIOT(n_chans=22, n_outputs=4, n_times=1000) # ... training code ... # Push to the Hub model.push_to_hub( repo_id="username/my-biot-model", commit_message="Initial model upload", )
Loading a model from the Hub:
from braindecode.models import BIOT # Load pretrained model model = BIOT.from_pretrained("username/my-biot-model") # Load with a different number of outputs (head is rebuilt automatically) model = BIOT.from_pretrained("username/my-biot-model", n_outputs=4)
Extracting features and replacing the head:
import torch x = torch.randn(1, model.n_chans, model.n_times) # Extract encoder features (consistent dict across all models) out = model(x, return_features=True) features = out["features"] # Replace the classification head model.reset_head(n_outputs=10)
Saving and restoring full configuration:
import json config = model.get_config() # all __init__ params with open("config.json", "w") as f: json.dump(config, f) model2 = BIOT.from_config(config) # reconstruct (no weights)
All model parameters (both EEG-specific and model-specific such as dropout rates, activation functions, number of filters) are automatically saved to the Hub and restored when loading.
See Loading and Adapting Pretrained Foundation Models for a complete tutorial.
Methods
- forward(x, *args, **kwargs)[source]#
Pass the input through the BIOT encoder, and then through the classification head.
- Parameters:
x (Tensor) – (batch_size, n_channels, n_times)
*args – The description is missing.
**kwargs – The description is missing.
- Returns:
Default:
torch.Tensorof shape(batch_size, n_outputs). Ifreturn_features=True:dictwith"features"(batch_size, emb_size)and"cls_token"(None). If legacyreturn_feature=True(init param):(out, emb)tuple (ignored whenreturn_features=True).- Return type:
torch.Tensor or tuple or dict
Examples using braindecode.models.InterpolatedBIOT#
Loading Pretrained Foundation Models on Arbitrary Channel Sets