braindecode.models.EEGModuleMixin#
- class braindecode.models.EEGModuleMixin(n_outputs=None, n_chans=None, chs_info=None, n_times=None, input_window_seconds=None, sfreq=None)[source]#
Mixin class for all EEG models in braindecode.
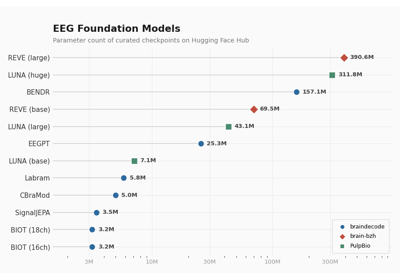
This class integrates with Hugging Face Hub when the
huggingface_hubpackage is installed, enabling models to be pushed to and loaded from the Hub usingpush_to_hub()andfrom_pretrained()methods.- Parameters:
n_outputs (
Optional[int]) – Number of outputs of the model. This is the number of classes in the case of classification.chs_info (list of dict) – Information about each individual EEG channel. This should be filled with
info["chs"]. Refer tomne.Infofor more details.n_times (
Optional[int]) – Number of time samples of the input window.input_window_seconds (
Optional[float]) – Length of the input window in seconds.sfreq (
Optional[float]) – Sampling frequency of the EEG recordings.
- Raises:
ValueError – If some input signal-related parameters are not specified: and can not be inferred.
Notes
If some input signal-related parameters are not specified, there will be an attempt to infer them from the other parameters.
Methods
- classmethod from_config(config)[source]#
Create a model instance from a configuration dict.
This is the inverse of
get_config(). Weights are not loaded – usefrom_pretrained()for that.- Parameters:
config (
dict) – Configuration dict as returned byget_config().- Returns:
A new model instance.
- Return type:
Examples
>>> import json >>> from braindecode.models import EEGNet >>> model = EEGNet(n_chans=22, n_times=1000, n_outputs=4, F1=16) >>> config = model.get_config() >>> # Reconstruct (without weights) >>> model2 = EEGNet.from_config(config) >>> model2.F1 16 >>> # Or from a JSON file >>> with open("config.json") as f: ... config = json.load(f) >>> model3 = EEGNet.from_config(config)
Added in version 1.4.
- get_config()[source]#
Return a JSON-serializable dict of all
__init__parameters.The returned dictionary can be saved to a JSON file and later used with
from_config()to reconstruct the model (without weights). It is also used internally bypush_to_hub()to persist the full model configuration.- Returns:
All
__init__parameters, JSON-serializable.type[nn.Module]parameters (e.g.activation) are encoded as importable dotted-path strings.- Return type:
Examples
>>> import json >>> from braindecode.models import EEGNet >>> model = EEGNet(n_chans=22, n_times=1000, n_outputs=4, F1=16) >>> config = model.get_config() >>> config["F1"] 16 >>> # Save to disk >>> with open("config.json", "w") as f: ... json.dump(config, f)
Added in version 1.4.
- get_torchinfo_statistics(col_names=('input_size', 'output_size', 'num_params', 'kernel_size'), row_settings=('var_names', 'depth'))[source]#
Generate table describing the model using torchinfo.summary.
- Parameters:
col_names (
Optional[Iterable[str]]) – Specify which columns to show in the output, see torchinfo for details, by default (“input_size”, “output_size”, “num_params”, “kernel_size”)row_settings (
Optional[Iterable[str]]) – Specify which features to show in a row, see torchinfo for details, by default (“var_names”, “depth”)
- Returns:
ModelStatistics generated by torchinfo.summary.
- Return type:
ModelStatistics
- reset_head(n_outputs)[source]#
Replace the classification head for a new number of outputs.
This is called automatically by
from_pretrained()when the user passes ann_outputsthat differs from the saved config. Override in subclasses that need a model-specific head structure.- Parameters:
n_outputs (int) – New number of output classes.
Examples
>>> from braindecode.models import BENDR >>> model = BENDR(n_chans=22, n_times=1000, n_outputs=4) >>> model.reset_head(10) >>> model.n_outputs 10
Added in version 1.4.
- to_dense_prediction_model(axis=(2, 3))[source]#
Transform a sequential model with strides to a model that outputs.
dense predictions by removing the strides and instead inserting dilations. Modifies model in-place.
- Parameters:
axis (
tuple[int,...] |int) – Axis to transform (in terms of intermediate output axes) can either be 2, 3, or (2,3).- Return type:
Notes
Does not yet work correctly for average pooling. Prior to version 0.1.7, there had been a bug that could move strides backwards one layer.
Examples using braindecode.models.EEGModuleMixin#
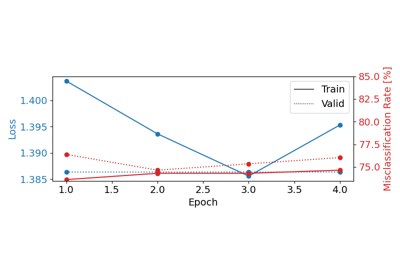
Cleaning EEG Data with EEGPrep for Trialwise Decoding
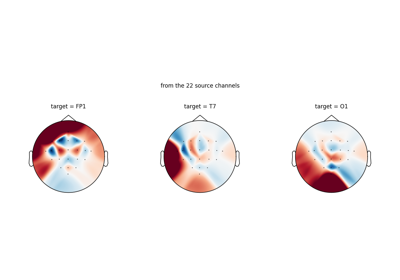
Loading Pretrained Foundation Models on Arbitrary Channel Sets
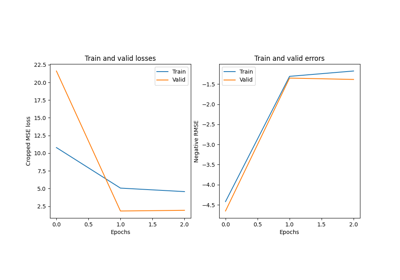
Convolutional neural network regression model on fake data.
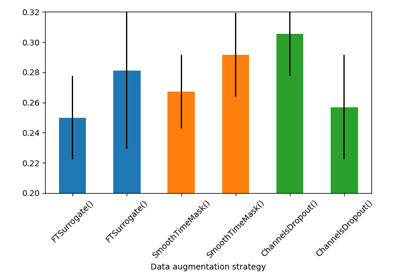
Searching the best data augmentation on BCIC IV 2a Dataset
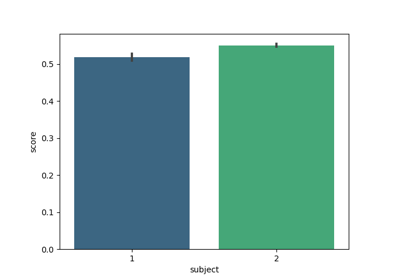
Cross-session motor imagery with deep learning EEGNet v4 model
Self-supervised learning on EEG with relative positioning

Sleep staging on the Sleep Physionet dataset using Chambon2018 network

Sleep staging on the Sleep Physionet dataset using Eldele2021
Sleep staging on the Sleep Physionet dataset using U-Sleep network