Note
Go to the end to download the full example code
Fingers flexion cropped decoding on BCIC IV 4 ECoG Dataset#
This tutorial shows you how to train and test deep learning models with Braindecode on ECoG BCI IV competition dataset 4 using cropped mode. For this dataset we will predict 5 regression targets corresponding to flexion of each finger. The targets were recorded as a time series (each 25 Hz), so this tutorial is an example of time series target prediction.
# Authors: Maciej Sliwowski <maciek.sliwowski@gmail.com>
# Mohammed Fattouh <mo.fattouh@gmail.com>
#
# License: BSD (3-clause)
Loading and preparing the dataset#
Loading#
First, we load the data. In this tutorial, we use the functionality of braindecode to load BCI IV competition dataset 4. The dataset is available as a part of ECoG library: https://searchworks.stanford.edu/view/zk881ps0522
The dataset contains ECoG signal and time series of 5 targets corresponding to each finger flexion. This is different than standard decoding setup for EEG with multiple trials and usually one target per trial. Here, fingers flexions change in time and are recorded with sampling frequency equals to 25 Hz.
If this dataset is used please cite [1].
[1] Miller, Kai J. “A library of human electrocorticographic data and analyses. “Nature human behaviour 3, no. 11 (2019): 1225-1235. https://doi.org/10.1038/s41562-019-0678-3
import copy
import numpy as np
import sklearn
from mne import set_log_level
from braindecode.datasets import BCICompetitionIVDataset4
subject_id = 1
dataset = BCICompetitionIVDataset4(subject_ids=[subject_id])
Split dataset into train and test#
We can easily split the dataset using additional info stored in the
description attribute, in this case session column. We select train dataset
for training and validation and test for final evaluation.
Preprocessing#
Now we apply preprocessing like bandpass filtering to our dataset. You can either apply functions provided by mne.Raw or mne.Epochs or apply your own functions, either to the MNE object or the underlying numpy array.
Note
Preprocessing steps are taken from a standard EEG processing pipeline. The only change is the cutoff frequency of the filter. For a proper ECoG decoding other preprocessing steps may be needed.
Note
These prepocessings are now directly applied to the loaded data, and not on-the-fly applied as transformations in PyTorch-libraries like torchvision.
from braindecode.preprocessing import (Preprocessor,
exponential_moving_standardize,
preprocess)
low_cut_hz = 1. # low cut frequency for filtering
high_cut_hz = 200. # high cut frequency for filtering, for ECoG higher than for EEG
# Parameters for exponential moving standardization
factor_new = 1e-3
init_block_size = 1000
We select only first 30 seconds from the training dataset to limit time and memory to run this example. We split training dataset into train and validation (only 6 seconds). To obtain full results whole datasets should be used.
valid_set = preprocess(copy.deepcopy(train_set),
[Preprocessor('crop', tmin=24, tmax=30)], n_jobs=-1)
preprocess(train_set, [Preprocessor('crop', tmin=0, tmax=24)], n_jobs=-1)
preprocess(test_set, [Preprocessor('crop', tmin=0, tmax=24)], n_jobs=-1)
<braindecode.datasets.base.BaseConcatDataset object at 0x7f4542b6bbb0>
In time series targets setup, targets variables are stored in mne.Raw object as channels of type misc. Thus those channels have to be selected for further processing. However, many mne functions ignore misc channels and perform operations only on data channels (see https://mne.tools/stable/glossary.html#term-data-channels).
preprocessors = [
# TODO: ensure that misc is not removed
Preprocessor('pick_types', ecog=True, misc=True),
Preprocessor(lambda x: x / 1e6, picks='ecog'), # Convert from V to uV
Preprocessor('filter', l_freq=low_cut_hz, h_freq=high_cut_hz), # Bandpass filter
Preprocessor(exponential_moving_standardize, # Exponential moving standardization
factor_new=factor_new, init_block_size=init_block_size, picks='ecog')
]
# Transform the data
preprocess(train_set, preprocessors)
preprocess(valid_set, preprocessors)
preprocess(test_set, preprocessors)
# Extract sampling frequency, check that they are same in all datasets
sfreq = train_set.datasets[0].raw.info['sfreq']
assert all([ds.raw.info['sfreq'] == sfreq for ds in train_set.datasets])
# Extract target sampling frequency
target_sfreq = train_set.datasets[0].raw.info['temp']['target_sfreq']
/home/runner/work/braindecode/braindecode/braindecode/preprocessing/preprocess.py:55: UserWarning: Preprocessing choices with lambda functions cannot be saved.
warn('Preprocessing choices with lambda functions cannot be saved.')
Create model#
In contrast to trialwise decoding, we first have to create the model before we can cut the dataset into windows. This is because we need to know the receptive field of the network to know how large the window stride should be.
We first choose the compute/input window size that will be fed to the network during training This has to be larger than the networks receptive field size and can otherwise be chosen for computational efficiency (see explanations in the beginning of this tutorial). Here we choose 1000 samples, which is 1 second for the 1000 Hz sampling rate.
input_window_samples = 1000
Now we create the deep learning model! Braindecode comes with some predefined convolutional neural network architectures for raw time-domain EEG. Here, we use the shallow ConvNet model from Deep learning with convolutional neural networks for EEG decoding and visualization. These models are pure PyTorch deep learning models, therefore to use your own model, it just has to be a normal PyTorch nn.Module.
import torch
from braindecode.models import ShallowFBCSPNet
from braindecode.util import set_random_seeds
cuda = torch.cuda.is_available() # check if GPU is available, if True chooses to use it
device = 'cuda' if cuda else 'cpu'
if cuda:
torch.backends.cudnn.benchmark = True
# Set random seed to be able to roughly reproduce results
# Note that with cudnn benchmark set to True, GPU indeterminism
# may still make results substantially different between runs.
# To obtain more consistent results at the cost of increased computation time,
# you can set `cudnn_benchmark=False` in `set_random_seeds`
# or remove `torch.backends.cudnn.benchmark = True`
seed = 20200220
set_random_seeds(seed=seed, cuda=cuda)
n_classes = 1
# Extract number of chans and time steps from dataset
n_chans = train_set[0][0].shape[0] - 5
model = ShallowFBCSPNet(
n_chans,
n_classes,
final_conv_length=2,
add_log_softmax=False,
)
# Send model to GPU
if cuda:
model.cuda()
from braindecode.models import get_output_shape, to_dense_prediction_model
to_dense_prediction_model(model)
/home/runner/.local/lib/python3.10/site-packages/sklearn/utils/deprecation.py:86: FutureWarning: Function to_dense_prediction_model is deprecated; will be removed in version 1.0. Use EEGModuleMixin.to_dense_prediction_model method directly on the model object.
warnings.warn(msg, category=FutureWarning)
To know the models’ receptive field, we calculate the shape of model output for a dummy input.
/home/runner/.local/lib/python3.10/site-packages/sklearn/utils/deprecation.py:86: FutureWarning: Function get_output_shape is deprecated; will be removed in version 1.0. Use EEGModuleMixin.get_output_shape method directly on the model object.
warnings.warn(msg, category=FutureWarning)
Cut Compute Windows#
from braindecode.preprocessing import create_fixed_length_windows
# Create windows using braindecode function for this. It needs parameters to define how
# trials should be used.
train_set = create_fixed_length_windows(
train_set,
start_offset_samples=0,
stop_offset_samples=None,
window_size_samples=input_window_samples,
window_stride_samples=n_preds_per_input,
drop_last_window=False,
targets_from='channels',
last_target_only=False,
preload=False
)
valid_set = create_fixed_length_windows(
valid_set,
start_offset_samples=0,
stop_offset_samples=None,
window_size_samples=input_window_samples,
window_stride_samples=n_preds_per_input,
drop_last_window=False,
targets_from='channels',
last_target_only=False,
preload=False
)
test_set = create_fixed_length_windows(
test_set,
start_offset_samples=0,
stop_offset_samples=None,
window_size_samples=input_window_samples,
window_stride_samples=n_preds_per_input,
drop_last_window=False,
targets_from='channels',
last_target_only=False,
preload=False
)
We select only the thumb’s finger flexion to create one model per finger.
Note
Methods to predict all 5 fingers flexion with the same model may be considered as well. We encourage you to find your own way to use braindecode models to predict fingers flexions.
Training#
In difference to trialwise decoding, we now should supply
cropped=True to the EEGClassifier, and CroppedLoss as the
criterion, as well as criterion__loss_function as the loss function
applied to the meaned predictions.
Note
In this tutorial, we use some default parameters that we have found to work well for EEG motor decoding, however we strongly encourage you to perform your own hyperparameter optimization using cross validation on your training data.
from skorch.callbacks import LRScheduler
from skorch.helper import predefined_split
from braindecode import EEGRegressor
from braindecode.training import CroppedTimeSeriesEpochScoring, TimeSeriesLoss
# These values we found good for shallow network for EEG MI decoding:
lr = 0.0625 * 0.01
weight_decay = 0
batch_size = 27 # only 27 examples in train set, otherwise set to 64
n_epochs = 8
regressor = EEGRegressor(
model,
cropped=True,
aggregate_predictions=False,
criterion=TimeSeriesLoss,
criterion__loss_function=torch.nn.functional.mse_loss,
optimizer=torch.optim.AdamW,
train_split=predefined_split(valid_set),
optimizer__lr=lr,
optimizer__weight_decay=weight_decay,
iterator_train__shuffle=True,
batch_size=batch_size,
callbacks=[
("lr_scheduler", LRScheduler('CosineAnnealingLR', T_max=n_epochs - 1)),
('r2_train', CroppedTimeSeriesEpochScoring(sklearn.metrics.r2_score,
lower_is_better=False,
on_train=True,
name='r2_train')
),
('r2_valid', CroppedTimeSeriesEpochScoring(sklearn.metrics.r2_score,
lower_is_better=False,
on_train=False,
name='r2_valid')
)
],
device=device,
)
set_log_level(verbose='WARNING')
Model training for a specified number of epochs. y is None as it is already supplied
in the dataset.
regressor.fit(train_set, y=None, epochs=n_epochs)
epoch r2_train r2_valid train_loss valid_loss lr dur
------- ---------- ---------- ------------ ------------ ------ ------
1 -16.9037 -8.3086 2.5378 20.7324 0.0006 0.4844
2 -13.7539 -7.1244 1.9596 18.0720 0.0006 0.4082
3 -12.7768 -6.7402 1.6787 17.1870 0.0005 0.4264
4 -11.7970 -6.3910 1.5638 16.4409 0.0004 0.4111
5 -11.2712 -6.1840 1.4237 16.0095 0.0002 0.4141
6 -10.5735 -5.9203 1.3549 15.4436 0.0001 0.4042
7 -9.8228 -5.6181 1.3132 14.7799 0.0000 0.4198
8 -9.0643 -5.3047 1.3316 14.0884 0.0000 0.4033
Obtaining predictions and targets for the test, train, and validation dataset
def pad_and_select_predictions(preds, y):
preds = np.pad(preds,
((0, 0), (0, 0), (y.shape[2] - preds.shape[2], 0)),
'constant',
constant_values=0)
mask = ~np.isnan(y[0, 0, :])
preds = np.squeeze(preds[..., mask], 0)
y = np.squeeze(y[..., mask], 0)
return y.T, preds.T
preds_train, y_train = regressor.predict_trials(train_set, return_targets=True)
preds_train, y_train = pad_and_select_predictions(preds_train, y_train)
preds_valid, y_valid = regressor.predict_trials(valid_set, return_targets=True)
preds_valid, y_valid = pad_and_select_predictions(preds_valid, y_valid)
preds_test, y_test = regressor.predict_trials(test_set, return_targets=True)
preds_test, y_test = pad_and_select_predictions(preds_test, y_test)
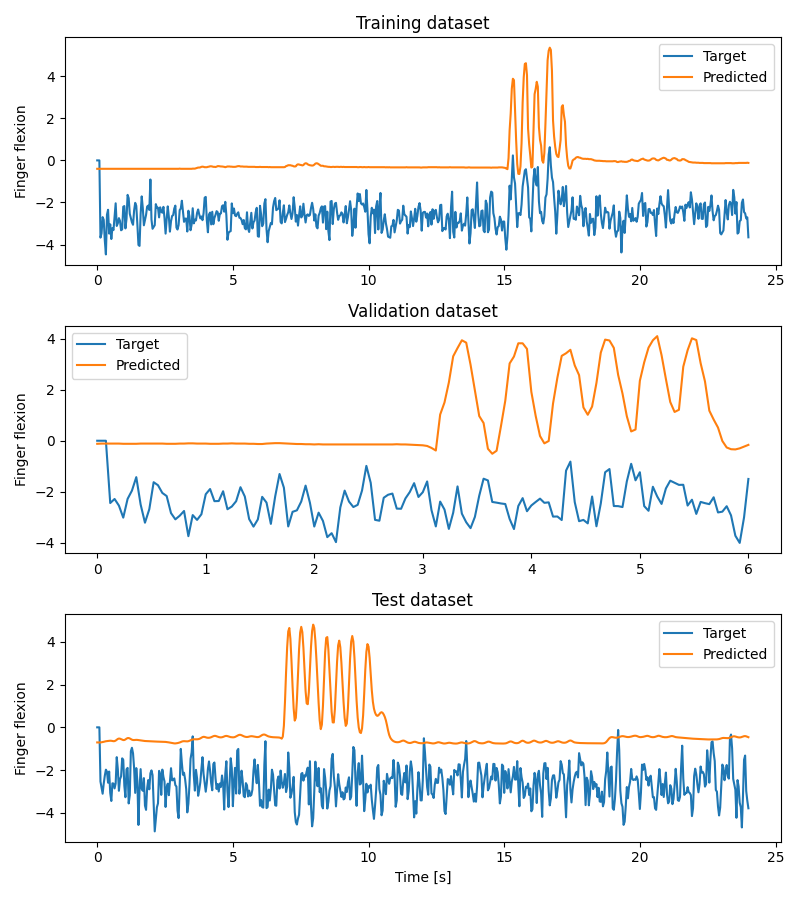
Plot Results#
We plot target and predicted finger flexion on training, validation, and test sets.
Note
The model is trained and validated on limited dataset (to decrease the time needed to run this example) which does not contain diverse dataset in terms of fingers flexions and may cause overfitting. To obtain better results use whole dataset as well as improve the decoding pipeline which may be not optimal for ECoG.
import matplotlib.pyplot as plt
import pandas as pd
from matplotlib.lines import Line2D
fig, axes = plt.subplots(3, 1, figsize=(8, 9))
axes[0].set_title('Training dataset')
axes[0].plot(np.arange(y_train.shape[0]) / target_sfreq, y_train[:, 0], label='Target')
axes[0].plot(np.arange(preds_train.shape[0]) / target_sfreq, preds_train[:, 0],
label='Predicted')
axes[0].set_ylabel('Finger flexion')
axes[0].legend()
axes[1].set_title('Validation dataset')
axes[1].plot(np.arange(y_valid.shape[0]) / target_sfreq, y_valid[:, 0], label='Target')
axes[1].plot(np.arange(preds_valid.shape[0]) / target_sfreq, preds_valid[:, 0],
label='Predicted')
axes[1].set_ylabel('Finger flexion')
axes[1].legend()
axes[2].set_title('Test dataset')
axes[2].plot(np.arange(y_test.shape[0]) / target_sfreq, y_test[:, 0], label='Target')
axes[2].plot(np.arange(preds_test.shape[0]) / target_sfreq, preds_test[:, 0], label='Predicted')
axes[2].set_xlabel('Time [s]')
axes[2].set_ylabel('Finger flexion')
axes[2].legend()
plt.tight_layout()
We can compute correlation coefficients for each finger
corr_coeffs = []
for dim in range(y_test.shape[1]):
corr_coeffs.append(
np.corrcoef(preds_test[:, dim], y_test[:, dim])[0, 1]
)
print('Correlation coefficient for each dimension: ', np.round(corr_coeffs, 2))
Correlation coefficient for each dimension: [-0.1]
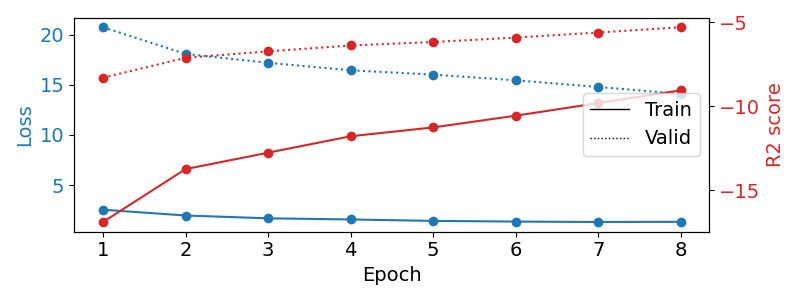
Now we use the history stored by Skorch throughout training to plot accuracy and loss curves. Extract loss and accuracy values for plotting from history object
results_columns = ['train_loss', 'valid_loss', 'r2_train', 'r2_valid']
df = pd.DataFrame(regressor.history[:, results_columns], columns=results_columns,
index=regressor.history[:, 'epoch'])
fig, ax1 = plt.subplots(figsize=(8, 3))
df.loc[:, ['train_loss', 'valid_loss']].plot(
ax=ax1, style=['-', ':'], marker='o', color='tab:blue', legend=False, fontsize=14)
ax1.tick_params(axis='y', labelcolor='tab:blue', labelsize=14)
ax1.set_ylabel("Loss", color='tab:blue', fontsize=14)
ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
df.loc[:, ['r2_train', 'r2_valid']].plot(
ax=ax2, style=['-', ':'], marker='o', color='tab:red', legend=False)
ax2.tick_params(axis='y', labelcolor='tab:red', labelsize=14)
ax2.set_ylabel("R2 score", color='tab:red', fontsize=14)
ax1.set_xlabel("Epoch", fontsize=14)
# where some data has already been plotted to ax
handles = []
handles.append(Line2D([0], [0], color='black', linewidth=1, linestyle='-',
label='Train'))
handles.append(Line2D([0], [0], color='black', linewidth=1, linestyle=':',
label='Valid'))
plt.legend(handles, [h.get_label() for h in handles], fontsize=14, loc='center right')
plt.tight_layout()
Total running time of the script: (1 minutes 24.674 seconds)
Estimated memory usage: 656 MB