Models Categorization#
Given the brain-decoding framework from the previous page, we define our neural networks, denoted \(f\), as a composition of sequential transformations:
where each \(f_\ell\) is a specific \(\ell\) layer in the neural network, focusing mostly of time in learning the mapping \(f_{\mathrm{method}} : \mathcal{X} \to \mathcal{Y}\) on the training data, with parameters \(\theta \in \Theta\). How these core \(\ell\) sequence transformations are structured and combined defines the overall focus and strength of the models.
Here, we categorize the main families of brain decoding models based on their core components and design philosophies. The categories are not mutually exclusive, but an indication of what governs that neural network model; many models blend elements from multiple families to leverage their combined strengths. Beginning directly, the categories are nine: Convolution, Filterbank, Interpretability, Recurrent, Attention/Transformer, Symmetric Positive-Definite, Graph Neural Network, Channel and Foundation Model.
At the moment, not all the categories are implemented, validated, and tested, but there are some that are noteworthy for introducing or popularizing concepts or layer designs that can take decoding further.
The convolutional layer appears as the core primitive across most architectures. This is because convolutions are filtering operations, such as band-pass filters, useful and needed to extract local features from brain signals. More details about each categories can be found in the respective sections below.
Convolution

Applies temporal and/or spatial convolutions to extract local features from brain signals.
Filterbank

Decomposes signals into multiple bands (learned or fixed) to capture frequency-specific information.
Interpretability
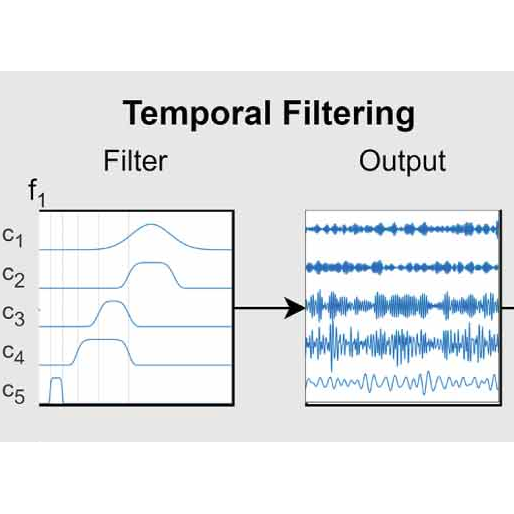
Architectures with inherently interpretable layers allow direct neuroscientific validation of learned features.
Recurrent
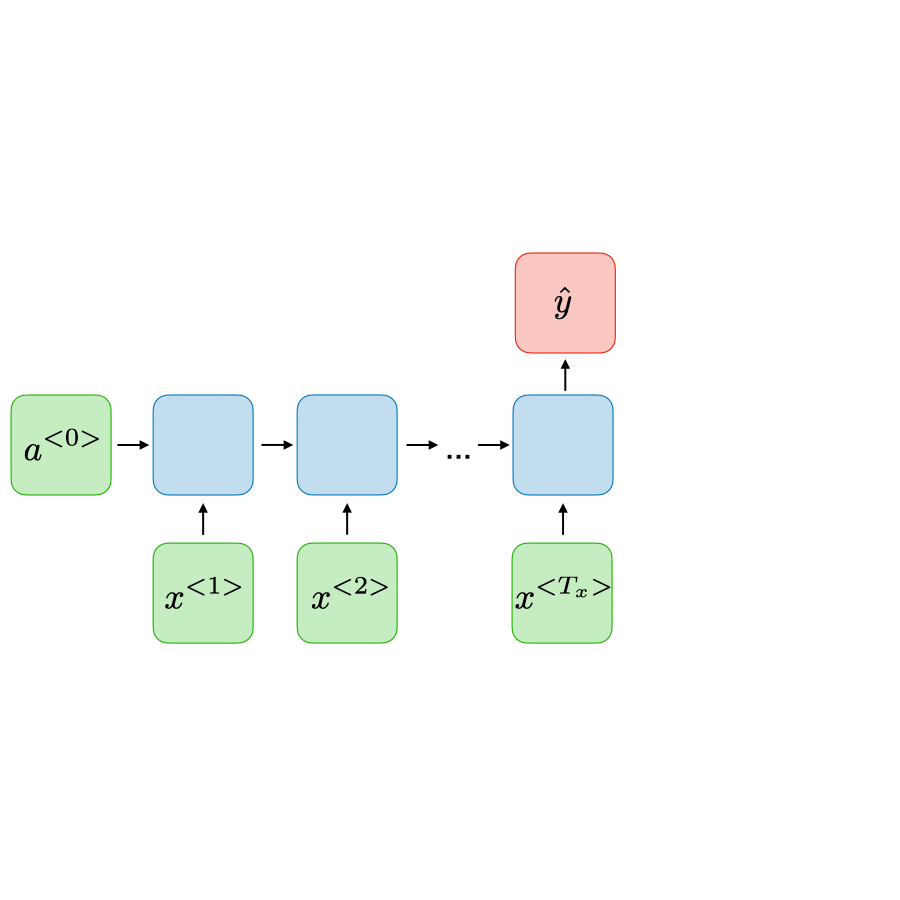
Models temporal dependencies via recurrent units or TCNs with dilations.
Attention/Transformer
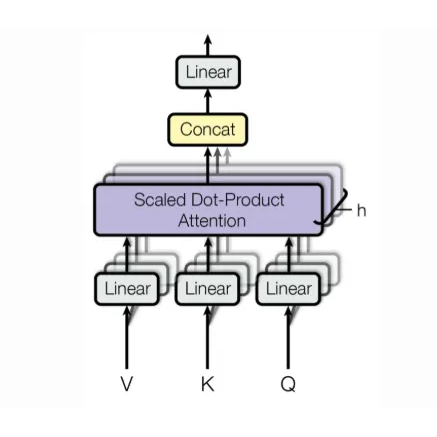
Uses attention mechanisms for feature focusing. Can be trained effectively without self-supervised pre-training.
SPD spd_learn

Learns on covariance/connectivity as SPD matrices using BiMap/ReEig/LogEig layers.
Graph Neural Network
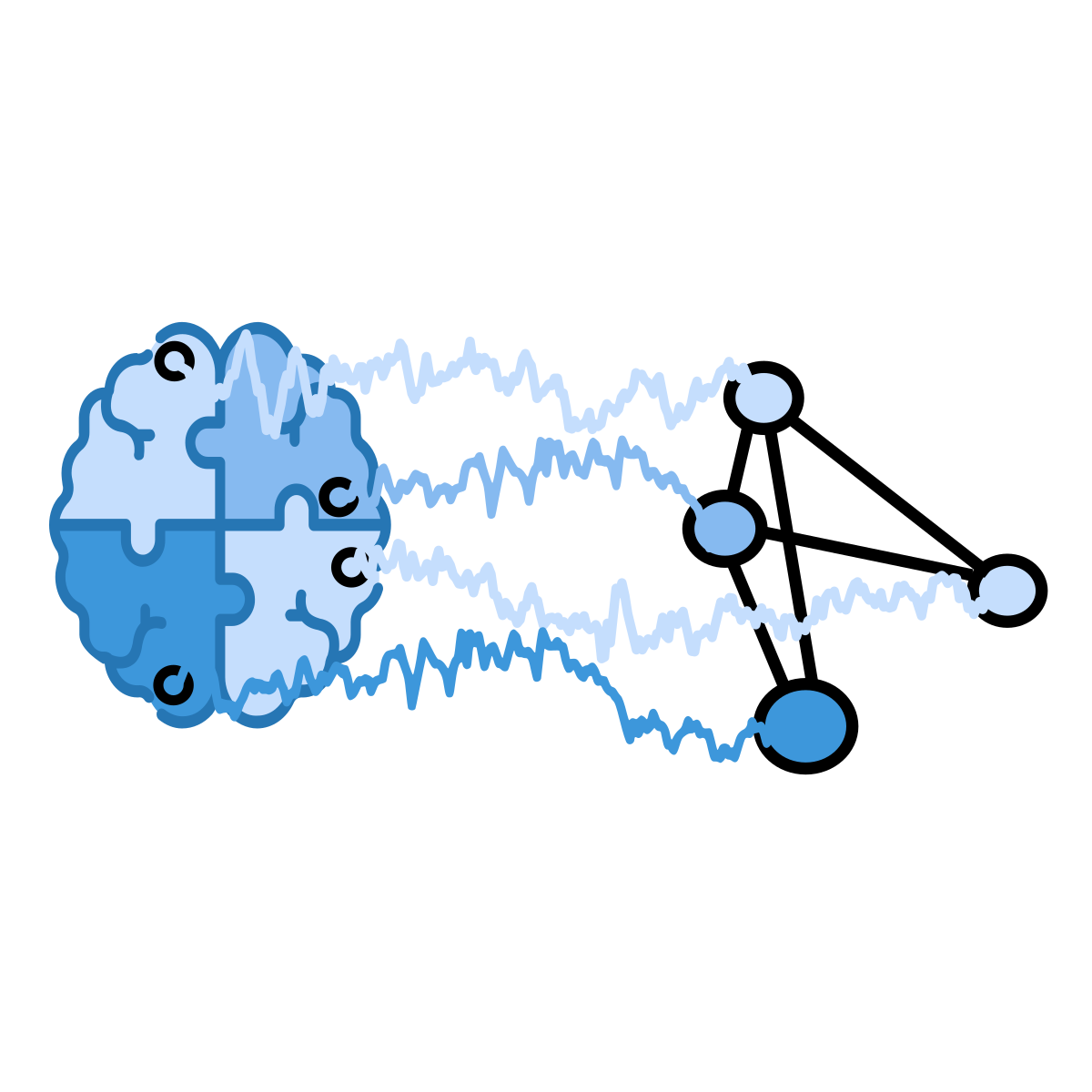
Treats channels/regions as nodes with learned/static edges to model connectivity.
Channel
Usage montage information with spatial filtering / channel / hemisphere / brain region selection strategies.
Foundation Model

Large-scale foundation model layers require self-supervised pre-training to work effectively.
Across most architectures, the earliest stages are convolutional (Convolution), reflecting the brain time series’s noisy, locally structured nature. These layers apply temporal and/or spatial convolutions—often depthwise-separable as in EEGNet, per-channel or across channel groups to extract robust local features.
EEGNet,ShallowFBCSPNet,EEGNeX, andEEGInceptionERPFilterbank-style models (Filterbank) explicitly decompose signals into multiple bands before (or while) learning, echoing the classic FBCSP pipeline; examples include
FBCNetandFBMSNet(11, 12).Interpretability-by-design (Interpretability) architectures expose physiologically meaningful primitives (e.g., band-pass/sinc filters, variance or connectivity features), enabling direct neuroscientific inspection; see
SincShallowNetandEEGMiner(13, 14).In the recurrent family (Recurrent), many modern EEG models actually rely on temporal convolutional networks (TCNs) with dilations to grow the receptive field, rather than explicit recurrence (15), such as
BDTCN.In contrast, several methods employ attention/transformer modules (Attention/Transformer) to capture longer-range dependencies efficiently, e.g.,
EEGConformer,CTNet,ATCNet,AttentionBaseNet, andEEGPT(16, 17, 18).SPD / Riemannian (SPD) methods operate on covariance (or connectivity) matrices as points on the SPD manifold, combining layers such as BiMap, ReEig, and LogEig; deep SPD networks and Riemannian classifiers motivate this family (19). Available via the spd_learn library (20).
Graph neural networks (Graph Neural Network) treat channels/regions as nodes with learned (static or dynamic) edges to model functional connectivity explicitly; e.g.,
DGCNN(21), more common in the emotion and epileptic decoding (22).Channel-domain robustness (Channel) techniques target variability in electrode layouts by learning montage-agnostic or channel-selective layers (e.g., dynamic spatial filtering, differentiable channel re-ordering); these strategies improve cross-setup generalization
SignalJEPA(23, 24).Foundation Model / Transformer (Foundation Model) approaches pretrain attention-based encoders on diverse biosignals and fine-tune for EEG tasks; e.g.,
BIOT(25),Labram(26), andEEGPT(27). These typically need a heavily self-supervised pre-training before decoding.
We are continually expanding this collection and welcome contributions! If you have implemented a model relevant to EEG, ECoG, or MEG analysis, consider adding it to Braindecode.
Submit a new model#
Want to contribute a new model to Braindecode? Great! You can propose a new model by opening an issue (please include a link to the relevant publication or description) or, even better, directly submit your implementation via a pull request. We appreciate your contributions to expanding the library!