 Symmetric Positive-Definite#
Symmetric Positive-Definite#
Symmetric Positive-Definite (SPD)
SPD / Riemannian methods operate on covariance (or connectivity) matrices as points on the SPD manifold, combining layers such as BiMap, ReEig, and LogEig. These models are available through the spd_learn library (20), a pure-PyTorch library for geometric deep learning on SPD matrices designed for neural decoding.
Installation#
pip install spd-learn
Available models#
Model |
Description |
|---|---|
Foundational architecture for deep learning on SPD manifolds; performs dimension reduction while preserving SPD structure using BiMap, ReEig, and LogEig layers (19). |
|
Specialized for brain-computer interface applications; combines covariance estimation with SPD network layers. |
|
Tangent Space Mapping Network that integrates convolutional processing with Riemannian geometry. |
|
SPDNet variant incorporating Tensor Common Spatial Patterns for multi-band EEG feature extraction. |
|
Leverages instantaneous phase information from analytic signals using Hilbert transforms. |
|
Gabor Riemann EEGNet combining Gabor wavelets with Riemannian geometry for robust EEG decoding. |
|
Matrix Attention network for SPD manifold learning. |
Citation#
If you use the SPD models, please cite the spd_learn library:
@article{aristimunha2026spd,
title={SPD Learn: A Geometric Deep Learning Python Library for Neural
Decoding Through Trivialization},
author={Aristimunha, Bruno and Ju, Ce and Collas, Antoine and
Bouchard, Florent and Mian, Ammar and Thirion, Bertrand and
Chevallier, Sylvain and Kobler, Reinmar},
journal={arXiv preprint arXiv:2602.22895},
year={2026}
}
LitMap#
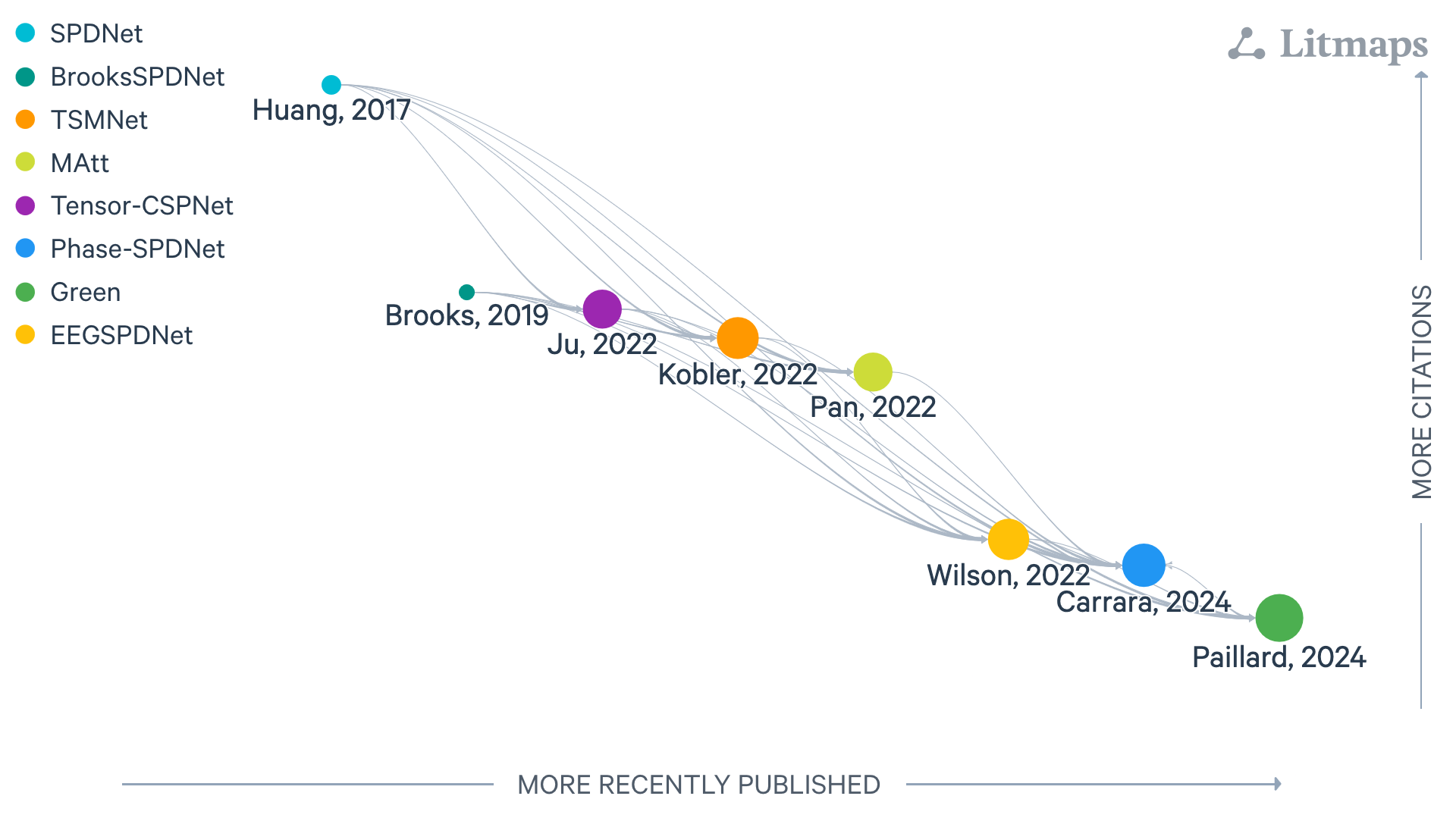
Figure: LitMap with symmetric positive-definite layers, last updated 26/08/2025. Each node is a paper; rightward means more recently published, upward more cited, and links show amount of citation with logaritm scale.#